Practical Biology
A collection of experiments that demonstrate biological concepts and processes.


Observing earthworm locomotion

Practical Work for Learning

Published experiments
Investigating effect of temperature on the activity of lipase, class practical.
Phenolphthalein is an indicator that is pink in alkaline solutions of about pH10. When the pH drops below pH 8.3 phenolphthalein goes colourless. Here, an alkaline solution of milk, lipase and phenolphthalein will change from pink to colourless as the fat in milk is broken down to form fatty acids (and glycerol ) thus reducing the pH to below 8.3. The time taken for this reaction to occur is affected by temperature .
Lesson organisation
This investigation could be carried out as a demonstration at two different temperatures, or in a group of at least 5 students with each student working at a different temperature. This would allow students to collect repeat data at their allocated temperature. Or it could be an investigation carried out by one student.
Apparatus and Chemicals
For each group of students:.
Test tube rack
Measuring cylinder (or syringe), 10 cm 3 , 2
Beaker, 100 cm 3 , 2 (for milk and sodium carbonate solution)
Beaker, 250 cm 3 , 2 (to act as water baths for temperatures below room temperature)
For each temperature: Thermometer
Syringe, 2 cm 3
Stop clock/stopwatch
For the class – set up by technician/ teacher:
Milk, full-fat or semi-skimmed, 5 cm 3 per student per temperature assessed
Phenolphthalein in a dropper bottle ( Note 2 )
5% lipase solution, 1 cm 3 per student per temperature assessed
Sodium carbonate solution, 0.05 mol dm – 3 , 7 cm 3 per student per temperature assessed
Electric hot water baths set to a range of temperatures, each containing a thermometer, a test-tube rack and a beaker of lipase solution.
Health & Safety and Technical notes
Sodium carbonate solution, 0.05 M. Make with 5.2 g of anhydrous solid, or 14.2 g of washing soda per litre of water. See CLEAPSS Hazcard; it is an IRRITANT at concentrations over 1.8 M.
Ethanol (IDA) in the phenolphthalein indicator is described as HIGHLY FLAMMABLE on the CLEAPSS Hazcard (flash point 13 °C) and HARMFUL (because of presence of methanol).
Glassware is breakable.
Electric water baths should be safety checked in accordance with your employer’s instructions.
Take care with thermometers and brief students how to react if they are broken.
Read our standard health & safety guidance
1 Lipase solution is best freshly made, but it will keep for a day or two in a refrigerator. Don’t try to study different temperatures on different days for the same investigation; the activity of the enzyme will change and it will not be a fair test.
2 Phenolphthalein is described as low hazard on CLEAPSS Hazcard. Refer to Recipe card (acid-base indicators): Dissolve 1 g in 600 cm 3 of IDA then make up to 1 litre with water. Label the bottle highly flammable. Suppliers of phenolphthalein solution may not use IDA; it also may be diluted. Follow any hazard warning on supplier’s bottles.
SAFETY: Keep the phenolphthalein solution away from sources of ignition.
Wear eye protection and quickly rinse any splashes of enzyme solution or sodium carbonate from the skin.
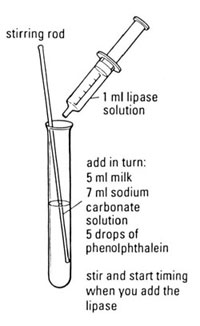
a Make up lipase solution and suitable quantities of the other solutions.
b Set up the water baths at a range of temperatures and put a beaker of lipase, containing a 2 cm 3 syringe into each water bath. Cover a range of temperatures up to around 60°C. An ice-bath will maintain a temperature of 0°C, until all the ice is melted.
Investigation
c Label a test tube with the temperature to be investigated.
d Add 5 drops of phenolphthalein to the test tube.
e Measure out 5 cm 3 of milk using a measuring cylinder (or syringe) and add this to the test tube.
f Measure out 7 cm 3 of sodium carbonate solution using another measuring cylinder (or syringe) and add this to the test tube. The solution should be pink.
g Place a thermometer in the test tube. Take care as the equipment could topple over.
h Place the test tube in a water bath and leave until the contents reach the same temperature as the water bath.
i Remove the thermometer from the test tube and replace it with a glass rod.
j Use the 2 cm 3 syringe to measure out 1 cm 3 of lipase from the beaker in the water bath for the temperature you are investigating.
k Add the lipase to the test tube and start the stopclock/ stopwatch.
l Stir the contents of the test tube until the solution loses its pink colour.
m Stop the clock/ watch and note the time in a suitable table of results.
Teaching notes
The quantities used should take approximately 4 minutes to change from pink to white at normal laboratory temperature. If this is not the case, change the concentration of enzyme to alter the speed of the reaction (more enzyme will reduce the time or increase the speed). Students will need to use the same volume at each temperature.
Digestion of fat produces fatty acids (and glycerol) that neutralise the alkali, sodium carbonate, thus lowering the pH and changing phenolphthalein from pink to colourless. You could use a pH probe or data logger, or another indicator.
You could add washing-up liquid to the solution (1 or 2 drops per 250 cm 3 ), to emulsify the fats which will provide a larger surface area for enzyme action. This will demonstrate the effect of bile salts. Or bile salts could be used.
Other factors to test:
- This protocol is based on a pH dependent result, so is not suitable for assessing the effect of different pHs on lipase.
- It would be possible to vary the concentration of the lipase and look at the effect of enzyme concentration on the breakdown of fat in milk.
- Different types of milk could be used Jersey, full cream, semi-skimmed and skimmed, to explore the effect on the reaction of changing fat concentration (substrate concentration).
Question 6 on the student question sheet opens the doors to a more extensive piece of research on this enzyme.
Health and safety checked, September 2008
Related experiments
Investigating an enzyme-controlled reaction: catalase and hydrogen peroxide concentration
Investigating effect of concentration on the activity of trypsin
Investigating the effect of pH on amylase activity
Stack Exchange Network
Stack Exchange network consists of 183 Q&A communities including Stack Overflow , the largest, most trusted online community for developers to learn, share their knowledge, and build their careers.
Q&A for work
Connect and share knowledge within a single location that is structured and easy to search.
How do bile salts affect lipase activity?
BACKGROUND : It is well known that bile salts are needed for emulsification of fats. It is then said that this increases the surface area for activity of pancreatic lipase, implying that bile salts make pancreatic lipase more effective. But it is also said that pancreatic lipase do need a colipase to overcome the inhibitory effect from bile salts .
QUESTION : So what does bile salts actually do? Do they increase or decrease lipase activity? How does colipase overcome it?
MY ATTEMPT : After some research , I found that bile salts above critical micelle concentration have inhibitory effect, but could not still comprehend how does the colipase makes lipase more efficient. Also if greater concentration is inhibitory then why does our body just secrete it in sufficient amounts? Isn't it be more efficient than to make a whole protein(colipase)?
- human-biology
- biochemistry
Let us first clear out some basic concepts regarding lipase, colipase and bile salts. The pancreatic lipase has an optimum pH range of about pH 8.0 ( Worthington ). This can be understood easily by Le Chatelier's principle : as the reaction moves forward, there is reduction of pH (due to formation of fatty acids). So, a basic pH would help the reaction in moving forward. See this diagram ( Worthington ):

It can also be understood on the basis of lipase's optimum pH, which lies between pH 8-9. See the graph (orange line) ( Chegg ):

This optimum pH is brought about by bile salts. Bile salts are slightly alkaline, with pH range of about 7-8 ( Britannica ). This helps lipase in catalysing its reaction. Bile salts also help lipase by increasing the surface area of fat droplets. Bile molecules have a hydrophobic and hydrophilic part. The hydrophobic part is attracted towards fat while the hydrophilic part is attracted towards water. This helps stabilize fat droplets by emulsification i.e. breaking the fat droplet(s) into smaller parts. This also increases the surface area of fat droplets.
Now, the matter gets complicated when we talk about concentration. Since bile molecules contain hydrophilic and hydrophobic ends, they tend to form micelles with increasing concentration (just like soap). As long as its concentration is below critical micelle concentration, it will carry on emulsification and increase lipase activity. But when its concentration becomes more than the critical micelle concentration, it will form micelles and not carry on emulsion. This significantly reduces lipase activity by not only decreasing surface area, but also reducing pH to slightly acidic ( Erlanson et al , 1973 ). This is where colipase helps. Colipase binds to the C-terminal, non-catalytic domain of lipase and stabilizes its active conformation and increases the hydrophobicity binding site ( Verger et al , 1999 ). Colipase strongly binds to the ester bond of the triglyceride molecule, by which its hydrolysis by lipase becomes easy ( Mansbach, 2011 ). Thus, colipase helps in overcoming the inhibitory effect or bile salts on lipase.
- $\begingroup$ Thanks. Why don't our body secrete bile salts in less than CMC. Won't that be more energy efficient than making a protein( in this case colipase)? $\endgroup$ – JM97 Commented Apr 26, 2017 at 10:50
- 1 $\begingroup$ It might be due to lack of regulation. All mechanisms I know about (CCK, secretin, enterohepatic circulation) only act as positive feedback loops. Also, gallbladder performs the job of concentrating bile, which doesn't seem to have any strong regulative mechanism either. $\endgroup$ – another 'Homo sapien' Commented Apr 26, 2017 at 11:09
- 1 $\begingroup$ The optimal pH of lipase probably has more to do with general acid/base catalysis than Le Chatelier. $\endgroup$ – canadianer Commented Apr 27, 2017 at 2:01
- $\begingroup$ @canadianer I just used Le Chatelier to give an idea of effect of pH on lipase. I'll check that point out too. Thanks for suggesting :) $\endgroup$ – another 'Homo sapien' Commented Apr 27, 2017 at 2:53
You must log in to answer this question.
Not the answer you're looking for browse other questions tagged human-biology biochemistry physiology enzymes digestion ..
- Featured on Meta
- Join Stack Overflow’s CEO and me for the first Stack IRL Community Event in...
- Bringing clarity to status tag usage on meta sites
Hot Network Questions
- What is the EPSG for Czechia (Czech) DMR 5G Lidar Data?
- Has anyone returned from space in a different vehicle from the one they went up in? And if so who was the first?
- Are all pass filters stable?
- Does Gödel’s incompleteness theorems imply the necessity of an infinite recursive hierarchy of “proofs”, and that any “proof” is relative?
- Would superhuman elites allow for more liberal governance?
- Can the planet Neptune be seen from Earth with binoculars?
- How would you read this time change with the given note equivalence?
- Can you move between an attack and the attack granted by Horde Breaker?
- Getting lost on a Circular Track
- Somebody used recommendation by an in-law – should I report it?
- How can I make this equation look better?
- Bathroom fan venting options
- On the convex cone of convex functions
- what is the purpose of long plastic sleeve tubes around connections in appliances
- Which volcano is more hazardous? Mount Rainier or Mount Hood?
- What is this phenomenon?
- Do US universities invite faculty applicants from outside the US for an interview?
- I want to be a observational astronomer, but have no idea where to start
- Is the warp core solely a power source or is it also the mechanism that produces the warp field?
- Switching x-axis and z-axis To appear instead of each other
- Finding nearest edge from face center
- `uname -r` shows kernel marked as `vmlinuz.old` in `/boot` partition
- What was used between these countertop sections?
- How cheap would rocket fuel have to be to make Mars colonization feasible (according to Musk)?
An official website of the United States government
The .gov means it’s official. Federal government websites often end in .gov or .mil. Before sharing sensitive information, make sure you’re on a federal government site.
The site is secure. The https:// ensures that you are connecting to the official website and that any information you provide is encrypted and transmitted securely.
- Publications
- Account settings
Preview improvements coming to the PMC website in October 2024. Learn More or Try it out now .
- Advanced Search
- Journal List
- Front Genet
Structure and Function of Pancreatic Lipase-Related Protein 2 and Its Relationship With Pathological States
Guoying zhu.
1 Department of Clinical Nutrition, Putuo People’s Hospital, School of Medicine, Tongji University, Shanghai, China
2 Department of Pediatrics Gastroenterology, School of Medicine, Washington University in St. Louis, St. Louis, MO, United States
Fengshang Zhu
3 Department of Gastroenterology, Tongji Hospital, School of Medicine, Tongji University, Shanghai, China
Dongping Huang
Changqing yang.
Pancreatic lipase is critical for the digestion and absorption of dietary fats. The most abundant lipolytic enzymes secreted by the pancreas are pancreatic triglyceride lipase (PTL or PNLIP) and its family members, pancreatic lipase-related protein 1 (PNLIPRP1or PLRP1) and pancreatic lipase-related protein 2 (PNLIPRP2 or PLRP2). Unlike the family’s other members, PNLIPRP2 plays an elemental role in lipid digestion, especially for newborns. Therefore, if genetic factors cause gene mutation, or other factors lead to non-expression, it may have an effect on fat digestion and absorption, on the susceptibility to pancreas and intestinal pathogens. In this review, we will summarize what is known about the structure and function of PNLIPRP2 and the levels of PNLIPRP2 and associated various pathological states.
Introduction
Effective digestion and absorption of dietary fats is important, which begins in the stomach by preduodenal lipase with a small amount of dietary triglyceride ( Miller and Lowe, 2008 ). Then, the partially digested emulsion particles empty into the duodenum, where it mixes with the pancreatic lipase secreted from the pancreas. In addition, the common bile duct from the gallbladder merges with the pancreatic duct, supplementing bile salts to the duodenum. The emulsion particles subsequently are hydrolyzed into liquid crystals containing monoglycerides, fatty acids, and cholesterol. Then, the digestion products are transformed by bile salts to the small intestine, taken up by enterocytes, or enter into lymphatic system ( Berton et al., 2009 ; Lindquist and Hernell, 2010 ). Hence, pancreatic lipase is critical for the digestion and absorption of dietary fats. The most abundant lipolytic enzymes secreted by the pancreas are pancreatic triglyceride lipase (PNLIP or PTL) and its family members, pancreatic lipase-related proteins 1 and 2 (PNLIPRP1/PLRP1 and PNLIPRP2/PLRP2). However, PNLIP is not expressed in neonatal humans and rodents; PNLIPRP1 has no lipase activity ( Roussel et al., 1998a ; Bakala et al., 2012 ). Therefore, PNLIPRP2 should have some different properties with PNLIP and plays a pivotal role in lipid digestion, especially for newborns. The purpose of this paper is to review the structure and function of PNLIPRP2 and the relationship between the expression of PNLIPRP2 and various pathological states.
The Biochemical Properties and Function of PNLIPRP2
PNLIPRP2 (GenBank Accession No. HSA149D17 ) is a member of the classic triglyceride lipase family. PNLIPRP1 and PNLIPRP2, respectively, have 68 and 65% homologous amino acid sequence with PNLIP ( Giller et al., 1992 ). The biggest difference in exons between PNLIP, PNLIPRP1, and PNLIPRP2 is that both PNLIP and PNLIPRP1 have 13 exons ( Figure 1 ), whereas PNLIPRP2 only has 12 exons, but contains a larger exon I (157 bp) ( Lowe, 2000 ). The larger exon I in PNLIPRP2 is even bigger than the sum of exon I and exon II of PNLIP or PNLIPRP1. The remaining exons are highly conserved ( Lowe, 2000 ). PNLIPRP2 is expressed in various tissues of different species. It was detected in the pancreas of animals including guinea pig, coypu, rabbit, horse, human, and rat; however, it was not expressed in the following species: ox, goat, sheep, dog, and cat ( De Caro et al., 2008 ). The length of human, rat, and mouse PNLIPRP2 mRNAs is 1,500 bp and its protein molecular weight is 53 kDa ( Lowe, 2000 ). PNLIPRP2 lipase has two domains: an N-terminal domain and a C-terminal domain from residues 18 to 353 and 354 to 466, respectively. The N-terminal domain consists of α/β hydrolase fold and the C-terminal has a β-sandwich structure ( Ollis et al., 1992 ). The length of the signal peptide may be 16 or 30 amino acids. It is a 30-amino acid signal peptide in the mouse and rat and a 16-amino acid signal peptide in human and coypu, but less 5′ sequence than the cDNAs isolated from the mouse or rat ( Payne et al., 1994 ; Lowe et al., 1998 ). Our colleagues’ findings suggest that PNLIPRP2’s expression can be detected before birth, at 15 weeks of gestation in humans and 17 days of gestation in rats and mice ( Yang et al., 2000 ).

The gene structure for PTL (PNLIP), PLRP1 (PNLIPRP1), and PLRP2 (PNLIPRP2). The exons are numbered with roman numerals. The number of nucleotides in each exon is given in each box. The biggest difference in exons is that PTL (PNLIP) and PLRP1 (PNLIPRP1) both have 13 exons, whereas PLRP2 (PNLIPRP2) only has 12 exons, but it contains a larger exon I (157 bp). PTL (PNLIP), pancreatic triglyceride lipase; PLRP1 (PNLIPRP1), pancreatic lipase-related protein 1; PLRP2 (PNLIPRP2), pancreatic lipase-related protein 2.
With the development of X-ray 3D structure technology, scholars can understand the interfacial recognition sites in the molecular structure of these enzymes and the conformational changes in the presence of lipids and amphiphiles ( Winkler et al., 1990 ; van Tilbeurgh et al., 1993 ). The active sites of many lipases are contained in the N-terminal domain and controlled by a so-called lid formed by a surface loop, β5 loop, and β9 loop. There is a catalytic triad, Ser152-His263-Asp176, at the bottom of this crevice ( Berton et al., 2007 ). This kind of lid makes lipase a special catalytic and interfacial activity at the water/oil interface, but shows low or no activity in a single water and oil phase. In the presence of lipase inhibitors, it undergoes conformational changes, then the solvents in the 3D structure of several lipases can be exposed to the active sites ( Egloff et al., 1995 ; Yang and Lowe, 2000 ; Eydoux et al., 2008 ). Nevertheless, the lid structure, β5 loop, and β9 loop act differently among various species. First is the difference between β5 loop and β9 loop, and second is whether the lid is an open conformation or not. Our previous studies show that the structural determinants of human PNLIPRP2 (HPNLIPRP2) lipase activity are the β5 loop and the lid domain, and the β9 loop inversely had smaller effects on activity ( Figure 2 ; Xiao and Lowe, 2015 ). In contrast, Eydoux et al. (2008) obtained different outcomes by making a crystal structure of HPNLIPRP2 in the absence of amphiphiles and found that the β9 loop is a crucial structural component involved in substrate binding. Dridi et al. (2013) confirmed the role of the β9 loop in the stabilization of the leaving acyl chain in lipolysis reaction on guinea pig PNLIPRP2 (GPNLIPRP2). In addition to the loop structure, another structural determinant of PNLIPRP2 lipase activity is the lid conformation. Eydoux et al. (2008) and our colleagues ( Xiao and Lowe, 2015 ) confirmed that the lid of HPNLIPRP2 adopts an open conformation in solution, contrary to what is observed with the human PNLIP. Therefore, the active site of HPNLIPRP2 might be directly accessible to a substrate. In contrast, the lid of rat PNLIPRP2 (RPNLIPRP2) lipase is in the closed conformation ( Mancheño et al., 2004 ; Valek et al., 2019 ). Many researchers, including our research group, illustrate the essential role of the lid in determining the substrate specificity and the mechanism of action of lipases ( Roussel et al., 1998b ; Yang et al., 2000 ; Berton et al., 2007 ; Eydoux et al., 2008 ), and the theory is that closed lid means interfacial activation. Consequently, RPNLIPRP2 lipase displayed interfacial activation at the water/oil interface, while HPNLIPRP2 lipase did not, which was considered a galactolipase. GPNLIPRP2 is the only PNLIPRP2 identified so far with a deletion in the lid domain ( Withers-Martinez et al., 1996 ), but it shows similar kinetic properties with HPNLIPRP2.

Structure of human PTL (PNLIP) and PLRP2 (PNLIPRP2) and the corresponding lid, β5, and β9 loops. (A) Superimposed α-carbon structure of PNLIP (blue) and PNLIPRP2 (yellow). (B) Surface structure of PNLIP showing the catalytic site cavity and the location of the lid domain (orange), β5 loop (blue), and β9 loop (yellow). (C) Superimposed β5 loops of PNLIP (blue) and PNLIPRP2 (yellow). The labeled amino acids are PNLIPRP2 residues. (D) Superimposed β9 loops of PNLIP (blue) and PNLIPRP2 (yellow). The labeled amino acids are PNLIPRP2 residues. PTL (PNLIP), pancreatic triglyceride lipase; PLRP2 (PNLIPRP2), pancreatic lipase-related protein 2.
Studying the relationship between the structure and function of lipase is of great significance for understanding the role of lipolysis and providing new targets for regulating lipase activity. Although three genes share most of the same structure but differ in their 3D structure and some amino acid sequences, their enzymatic properties are different among them.
Hydrolyzed Substrate
Substrate specificity is strongly based on the supramolecular organization of the lipid substrates present in oil-in-water emulsions, membranes, micelles, monolayers, or vesicles. PNLIPRP2 had a high activity on all phospholipid–bile salt micelles. They can modify the properties of lipid/water interfaces and promote the enzyme–micelle interaction, thus initiating the effective mass transfer between micelles and enzymes during lipolysis reaction ( Mateos-Diaz et al., 2018 ). PNLIPRP2 has a broader substrate specificity and can hydrolyze triglycerides, phospholipids, and galactolipids. PNLIP has no effect on activity against the phospholipid and galactolipid substrates, except triglycerides ( Lowe, 2002 ; Mancheño et al., 2004 ), and PNLIPRP1 shows no lipase activity against all known substrates ( Roussel et al., 1998 ).
Effects of Colipase and Bile Salts on Kinetic Properties
Neonatal and lactating infants express colipase and PNLIPRP2, but not PNLIP. PNLIP is inhibited by normal components of the duodenum, for example, bile acids, phospholipids, or dietary proteins, and colipase can reverse the inhibition of PNLIP ( Lowe, 1994 ; D’Agostino and Lowe, 2004 ; De Caro et al., 2004 ; Xiao et al., 2013 ). The activity of PNLIPRP2 varies greatly among species. HPNLIPRP2 is considered to be a galactolipase, which was inhibited by bile salts when against long-chain triglycerides, with poor activity against diglycerides ( Eydoux et al., 2007 ; Amara et al., 2009 ; Pang et al., 2011 ). Horse PNLIPRP2 is the same with HPNLIPRP2 ( Jayne et al., 2002b ). On the contrary, our research ( Xiao et al., 2011 ) found that HPNLIPRP2 had sufficient activity against long-chain triglycerides and diolein in the presence of bile salt micelles, in vitro , and depended on colipase. Under optimal conditions, the activity of mouse PNLIPRP2 (MPNLIPRP2) is about seven-fold greater than HPNLIPRP2 when against long-chain triglycerides and 10-fold higher than HPNLIPRP2 when against short- and medium-chain triglycerides. RPNLIPRP2 and MPNLIPRP2 have activity in the presence of bile salt micelles, and their activity can be increased by colipase ( Jennens and Lowe, 1995a , b ; D’Agostino and Lowe, 2004 ). MPNLIPRP2 had full activity in the presence of bovine serum albumin (BSA), whereas BSA completely inhibited MPNLIP except for the presence of colipase ( D’Agostino and Lowe, 2004 ). Why are the effects of colipase and bile salts different among these structurally enzymes? Lowe and Jayne ( Jayne et al., 2002a , b ; Johnson et al., 2013 ) suggested that colipase stimulates the activity of PNLIPRP2 by acting on the substrate rather than by anchoring PNLIPRP2 to the substrate interface as the colipase–PNLIP complex does. Therefore, PNLIPRP2 has activity with or without colipase and the degree of activity stimulated by colipase depended on the substrate and PNLIPRP2 species.
Function of PNLIPRP2
Pancreatic lipase is usually secreted by the pancreas and transferred to the duodenum to participate in the hydrolysis and digestion of fat, cholesterol esters, and fat-soluble vitamins ( Carrière et al., 1994 ). The temporal pattern of PNLIPRP2 mRNA expression confirmed by many experimental data suggests that PNLIPRP2 may play an important role in milk fat digestion in lactating mammals ( Li et al., 2007 ; Andersson et al., 2011 ). Sucking PNLIPRP2-deficient mice were found to have steatorrhea and fat malabsorption, and the undigested and partially digested triglycerides in feces were significantly increased, accompanied by a significant decrease in weight gain curve ( D’Agostino et al., 2002 ; Huggins et al., 2003 ; De Caro et al., 2004 ; Gilham et al., 2007 ). Intriguingly, as a presumed galactolipase, the main enzyme of HPNLIPRP2 was involved in the digestion of those common vegetable lipids in the gastrointestinal tract, but there is dissimilarity in various species ( Bourne et al., 1994 ; Carrière et al., 1998 ; Aloulou et al., 2006 ; Amara et al., 2010 ). It was detectable in the pancreas of both omnivorous and monogastric herbivorous animals, but not of carnivorous and ruminant herbivorous species, turkey, pigs, and ostrich ( De Caro et al., 2008 ). Galactolipids in the plant kingdom are much more abundant than triacylglycerols, which are ingested by galactolipase-PNLIPRP2. Hence, HPNLIPRP2 likely has some relationships with various races which have diverse component diets.
Between PNLIPRP2 Levels and Various Pathological States
Pnliprp2 levels and pancreatitis.
More and more lines of evidence show that the expression level of HPNLIPRP2 is related to chronic pancreatitis (CP). It was significantly lower in patients with chronic calcifying pancreatitis (CCP) than in the control group, and the ratio of HPNLIPRP2 to HPNLIP was 23.96% (W/W) and 28.3% (W/W) in CCP patients and controls, respectively ( Eydoux et al., 2006 ). On the contrary, Khatua et al. (2019) found that the expression level of PNLIPRP2 was elevated in fat necrosis and might regulate lipolysis and lipotoxic injury during pancreatitis. The possible explanation is that the secretion of lipase and the occurrence and development of pancreatitis are dynamic processes. Anyway, hPNLIPRP2 is abnormally expressed in subjects with pancreatitis. Intriguingly, more recent studies have shown that genetic variants in pancreatic lipases are associated with an increased risk of CP. One report showed that two brothers with PNLIP deficiency were found to be homozygous for missense mutation in PNLIP and associated with CP ( Behar et al., 2014 ). The PNLIPRP2 W358X (the same with p.W357X and p.W340X) SNP is also of particular interest since it is a common non-sense polymorphism and present in different ethnic groups at a high allele frequency from 0.3 to 0.5. The genetic polymorphism results in a truncated protein, premature truncation of about −24% of the gene product, lacking nearly the entire C-terminal domain of HPNLIPRP2, which is necessary for its stability, efficient secretion, and full activity ( Jennens and Lowe, 1995b ; Cao and Hegele, 2003 ). The experience of our research group ( Xiao et al., 2013 ) concluded that the aberrant folding of W358X mutant may cause chronic cellular stress in pancreatic exocrine cells and increase susceptibility to other metabolic stressors. However, Németh et al. (2018) found that the p.W358X truncation variant of HPNLIPRP2 is expressed poorly and has no significant effect on the risk of CP. As a result, it deserved further investigations or more data to elucidate the discrepancy.
PNLIPRP2 Levels and Other Pathological States
PNLIPRP2 was secreted not only from the pancreas but also from various tissues and cell types under certain conditions, such as cytotoxic T lymphocytes (CTL). It may play an auxiliary role in some types of cytotoxic T-cell-mediated lysis ( Alves et al., 2009 ). Rabbit PNLIPRP2 (also named GP-3) associated with the zymogen granule membranes was detected in enterocytes and Paneth cells ( Grusby et al., 1990 ; Wagner et al., 1994 ). In the rat hypothalamus, compared with the control group, it was downregulated during fasting (seven-fold) and upregulated (1.8-fold) during conditions of metabolic excess ( Rippe et al., 2007 ). Moreover, it was also regulated by a high-fat (HF) diet at the post-transcriptional level in C57BL/6J mice ( Birk et al., 2014 ). MPNLIRP2 is associated with the hydrolysis of hepatic retinyl esters for the utilization of vitamin A in the mouse liver ( De Caro et al., 2004 ; Reboul et al., 2006 ) and is responsible for the increased hepatic retinyl ester hydrolases in mice fed vitamin A-deficient diet ( Gao et al., 2019 ). Goat PNLIPRP2 (GoPNLIPRP2) might be regulated by the sexual hormones, because its expression in seminal plasma was significantly increased during the breeding season, parallel to the increase in the plasmatic levels of testosterone ( Sias et al., 2005 ). The low expression of lipases resulted in the delivery of undigested lipid components to the distal ileum, where their intracellular accumulation can lead to the generation of reactive oxygen species (ROS) oxidative stress and the inflammatory characteristics of necrotizing enterocolitis (NEC) ( Sodhi et al., 2018 ). All these data raised the possibility that PNLIPRP2 has other significant functions than just hydrolyzing dietary fats.
The structural parameters are responsible for the substrate specificity among these structural enzymes, and the degree of activity depends on the substrate and PNLIPRP2 species. These points remain, however, speculations and will deserve further structural studies to determine the conformational state of the PNLIPRP2 lid more precisely and also further investigations to elucidate the molecular mechanisms of PNLIPRP2 processing along with detailed analysis of the digestion products. It might pave the way for exploiting the different expressions and functions of PNLIPRP2 among species, different mutations of PNLIPRP2 among various races, and the relationship between PNLIPRP2 levels and various pathological states and for providing a new drug target to modulate lipase activity.
Author Contributions
DH and CY conceptualized and supervised the manuscript. GZ, QF, and FZ contributed to the drafting and editing of the manuscript. All authors contributed to the article and approved the submitted version.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Xunjun Xiao and Mark E. Lowe from the Department of Pediatric Gastroenterology, School of Medicine, Washington University, St. Louis, for selflessly providing their related published papers and guidance to this manuscript.
Funding. This work was supported by grants from the Shanghai Municipal Key Clinical Specialty (shslczdzk06801, China) to CY and Shanghai Medical Key Specialty Fund (ZK2019C16, China) to DH.
- Aloulou A., Rodriguez J. A., Fernandez S., van Oosterhout D., Puccinelli D., Carrière F. (2006). Exploring the specific features of interfacial enzymology based on lipase studies. Biochim. Biophys. Acta 1761 995–1013. 10.1016/j.bbalip.2006.06.009 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Alves B. N., Marshall K., Tamang D. L., Leong J., Redelman D., Elliott V., et al. (2009). Lipid-dependent cytotoxicity by the lipase PLRP2 and by PLRP2-positive cytotoxic T lymphocytes (CTLs). Cell Biochem. Funct . 27 296–308. 10.1002/cbf.1573 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Amara S., Barouh N., Lecomte J., Lafont D., Robert S., Villeneuve P., et al. (2010). Lipolysis of natural long chain and synthetic medium chain galactolipids by pancreatic lipase-related protein 2. Biochim. Biophys. Acta 1801 508–516. 10.1016/j.bbalip.2010.01.003 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Amara S., Lafont D., Fiorentino B., Boullanger P., Carrière F., De Caro A. (2009). Continuous measurement of galactolipid hydrolysis by pancreatic lipolytic enzymes using the pH-stat technique and a medium chain monogalactosyl diglyceride as substrate. Biochim. Biophys. Acta 1791 983–990. 10.1016/j.bbalip.2009.05.002 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Andersson E. L., Hernell O., Bläckberg L., Fält H., Lindquist S. (2011). BSSL and PLRP2: key enzymes for lipid digestion in the newborn examined using the Caco-2 cell line. J. Lipid Res. 52 1949–1956. 10.1194/jlr.M015685 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Bakala N., Goma J. C., Amara S., Dridi K., Jannin V., Carrière F. (2012). Understanding the lipid-digestion processes in the GI tract before designing lipid-based drug-delivery systems. Ther. Deliv. 3 105–124. 10.4155/tde.11.138 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Behar D. M., Basel-Vanagaite L., Glaser F., Kaplan M., Tzur S., Magal N., et al. (2014). Identification of a novel mutation in the PNLIP gene in two brothers with congenital pancreatic lipase deficiency. J. Lipid Res. 55 307–312. 10.1194/jlr.P041103 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Berton A., Sebban-Kreuzer C., Crenon I. (2007). Role of the structural domains in the functional properties of pancreatic lipase-related protein 2. FEBS J. 274 6011–6023. 10.1111/j.1742-4658.2007.06123.x [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Berton A., Sebban-Kreuzer C., Rouvellac S., Lopez C., Crenon I. (2009). Individual and combined action of pancreatic lipase and pancreatic lipase-related proteins 1 and 2 on native versus homogenized milk fat globules. Mol. Nutr. Food Res. 53 1592–1602. 10.1002/mnfr.200800563 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Birk R. Z., Rubio-Aliaga I., Boekschoten M. V., Danino H., Müller M., Daniel H. (2014). Differential regulation of pancreatic digestive enzymes during chronic high-fat diet-induced obesity in C57BL/6J mice. Br. J. Nutr. 112 154–161. 10.1017/S0007114514000816 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Bourne Y., Martinez C., Kerfelec B., Lombardo D., Chapus C., Cambillau C. (1994). Horse pancreatic lipase. The crystal structure refined at 2.3 A resolution. J. Mol. Biol. 238 709–732. 10.1006/jmbi.1994.1331 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Cao H., Hegele R. A. (2003). DNA polymorphisms of lipase related genes. J. Hum. Genet. 48 443–446. 10.1007/s10038-003-0051-1 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Carrière F., Thirstrup K., Hjorth S., Boel E. (1994). Cloning of the classical guinea pig pancreatic lipase and comparison with the lipase related protein 2. FEBS Lett. 338 63–68. 10.1016/0014-5793(94)80117-7 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Carrière F., Withers-Martinez C., van Tilbeurgh H., Roussel A., Cambillau C., Verger R. (1998). Structural basis for the substrate selectivity of pancreatic lipases and some related proteins. Biochim. Biophys. Acta 1376 417–432. 10.1016/s0304-4157(98)00016-1 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- D’Agostino D., Cordle R. A., Kullman J., Erlanson-Albertsson C., Muglia L. J., Lowe M. E. (2002). Decreased postnatal survival and altered body weight regulation in procolipase-deficient mice. J. Biol. Chem. 277 7170–7177. 10.1074/jbc.M108328200 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- D’Agostino D., Lowe M. E. (2004). Pancreatic lipase-related protein 2 is the major colipase-dependent pancreatic lipase in suckling mice. J.Nutr. 134 132–134. 10.1093/jn/134.1.132 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- De Caro J., Eydoux C., Chérif S., Lebrun R., Gargouri Y., Carrière F., et al. (2008). Occurrence of pancreatic lipase-related protein-2 in various species and its relationship with herbivore diet. Comp. Biochem. Physiol. B Biochem. Mol. Biol. 150 1–9. 10.1016/j.cbpb.2008.01.007 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- De Caro J., Sias B., Grandval P., Ferrato F., Halimi H., Carrière F., et al. (2004). Characterization of pancreatic lipase-related protein 2 isolated from human pancreatic juice. Biochim. Biophys. Acta 1701 89–99. 10.1016/j.bbapap.2004.06.005 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Dridi K., Amara S., Bezzine S., Rodriguez J. A., Carrière F., Gaussier H. (2013). Partial deletion of beta9 loop in pancreatic lipase-related protein 2 reduces enzyme activity with a larger effect on long acyl chain substrates. Biochim. Biophys. Acta 1831 1293–1301. 10.1016/j.bbalip.2013.04.010 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Egloff M. P., Marguet F., Buono G., Verger R., Cambillau C., van Tilbeurgh H. (1995). The 2.46 A resolution structure of the pancreatic lipase-colipase complex inhibited by a C11 alkyl phosphonate. Biochemistry 34 2751–2762. 10.1021/bi00009a003 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Eydoux C., Aloulou A., De Caro J., Grandval P., Laugier R., Carrière F., et al. (2006). Human pancreatic lipase-related protein 2: tissular localization along the digestive tract and quantification in pancreatic juice using a specific ELISA. Biochim. Biophys. Acta 1760 1497–1504. 10.1016/j.bbagen.2006.06.005 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Eydoux C., De Caro J., Ferrato F., Boullanger P., Lafont D., Laugier R., et al. (2007). Further biochemical characterization of human pancreatic lipase-related protein 2 expressed in yeast cells. J. Lipid Res. 48 1539–1549. 10.1194/jlr.M600486-JLR200 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Eydoux C., Spinelli S., Davis T. L., Walker J. R., Seitova A., Dhe-Paganon S., et al. (2008). Structure of human pancreatic lipase-related protein 2 with the lid in an open conformation. Biochemistry 47 9553–9564. 10.1021/bi8005576 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Gao Y., Lu W., Sun Q., Yang X., Liu J., Ge W., et al. (2019). Pancreatic lipase-related protein 2 is responsible for the increased hepatic retinyl ester hydrolase activity in vitamin A-deficient mice. FEBS J. 286 4232–4244. 10.1111/febs.14958 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Gilham D., Labonté E. D., Rojas J. C., Jandacek R. J., Howles P. N., Hui D. Y. (2007). Carboxyl ester lipase deficiency exacerbates dietary lipid absorption abnormalities and resistance to diet-induced obesity in pancreatic triglyceride lipase knockout mice. J. Biol. Chem. 282 24642–24649. 10.1074/jbc.M702530200 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Giller T., Buchwald P., Blum-Kaelin D., Hunziker W. (1992). Two novel human pancreatic lipase related proteins, hPLRP1 and hPLRP2. Differences in colipase dependence and in lipase activity. J. Biol. Chem. 267 16509–16516. [ PubMed ] [ Google Scholar ]
- Grusby M. J., Nabavi N., Wong H., Dick R. F., Bluestone J. A., Schotz M. C., et al. (1990). Cloning of an interleukin-4 inducible gene from cytotoxic T lymphocytes and its identification as a lipase. Cell 60 451–459. 10.1016/0092-8674(90)90596-7 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Huggins K. W., Camarota L. M., Howles P. N., Hui D. Y. (2003). Pancreatic triglyceride lipase deficiency minimally affects dietary fat absorption but dramatically decreases dietary cholesterol absorption in mice. J. Biol. Chem. 278 42899–42905. 10.1074/jbc.M303422200 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Jayne S., Kerfelec B., Foglizzo E., Chapus C., Crenon I. (2002a). High expression in adult horse of PLRP2 displaying a low phospholipase activity. Biochim. Biophys. Acta 1594 255–265. 10.1016/s0167-4838(01)00309-0 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Jayne S., Kerfelec B., Foglizzo E., Granon S., Hermoso J., Chapus C., et al. (2002b). Activation of horse PLRP2 by bile salts does not require colipase. Biochemistry 41 8422–8428. 10.1021/bi025867j [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Jennens M. L., Lowe M. E. (1995a). C-terminal domain of human pancreatic lipase is required for stability and maximal activity but not colipase reactivation. J. Lipid Res . 36 1029–1036. [ PubMed ] [ Google Scholar ]
- Jennens M. L., Lowe M. E. (1995b). Rat GP-3 is a pancreatic lipase with kinetic properties that differ from colipase-dependent pancreatic lipase. J. Lipid Res. 36 2374–2382. [ PubMed ] [ Google Scholar ]
- Johnson K., Ross L., Miller R., Xiao X., Lowe M. E. (2013). Pancreatic lipase-related protein 2 digests fats in human milk and formula in concert with gastric lipase and carboxyl ester lipase. Pediatr. Res. 74 127–132. 10.1038/pr.2013.90 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Khatua B., Trivedi R. N., Noel P., Patel K., Singh R., de Oliveira C., et al. (2019). Carboxyl Ester Lipase May Not Mediate Lipotoxic Injury during Severe Acute Pancreatitis. Am. J. Pathol. 189 1226–1240. 10.1016/j.ajpath.2019.02.015 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Li X., Lindquist S., Lowe M., Noppa L., Hernell O. (2007). Bile salt-stimulated lipase and pancreatic lipase-related protein 2 are the dominating lipases in neonatal fat digestion in mice and rats. Pediatr. Res. 62 537–541. 10.1203/PDR.0b013e3181559e75 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Lindquist S., Hernell O. (2010). Lipid digestion and absorption in early life: an update. Curr. Opin. Clin. Nutr. Metab. Care 13 314–320. 10.1097/MCO.0b013e328337bbf0 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Lowe M. E. (1994). Pancreatic triglyceride lipase and colipase: insights into dietary fat digestion. Gastroenterology 107 1524–1536. 10.1016/0016-5085(94)90559-2 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Lowe M. E. (2000). Properties and function of pancreatic lipase related protein 2. Biochimie 82 997–1004. 10.1016/s0300-9084(00)01184-6 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Lowe M. E. (2002). The triglyceride lipases of the pancreas. J. Lipid Res. 43 2007–2016. 10.1194/jlr.r200012-jlr200 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Lowe M. E., Kaplan M. H., Jackson-Grusby L., D’Agostino D., Grusby M. J. (1998). Decreased neonatal dietary fat absorption and T cell cytotoxicity in pancreatic lipase-related protein 2-deficient mice. J. Biol. Chem. 273 31215–31221. 10.1074/jbc.273.47.31215 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Mancheño J. M., Jayne S., Kerfelec B., Chapus C., Crenon I., Hermoso J. A. (2004). Crystallization of a proteolyzed form of the horse pancreatic lipase-related protein 2: structural basis for the specific detergent requirement. Acta Crystallogr.D Biol.Crystallogr. 60 2107–2109. 10.1107/S0907444904024229 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Mateos-Diaz E., Sutto-Ortiz P., Sahaka M., Byrne D., Gaussier H., Carrière F. (2018). IR spectroscopy analysis of pancreatic lipase-related protein 2 interaction with phospholipids: 2. Discriminative recognition of various micellar systems and characterization of PLRP2-DPPC-bile salt complexes. Chem. Phys. Lipids 211 66–76. 10.1016/j.hemphyslip.20.17.11.012 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Miller R., Lowe M. E. (2008). Carboxyl ester lipase from either mother’s milk or the pancreas is required for efficient dietary triglyceride digestion in suckling mice. J. Nutr . 138 927–930. 10.1093/jn/138.5.927 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Németh B. C., Pesei Z. G., Hegyi E., Szücs Á, Szentesi A., Hegyi P., et al. (2018). The common truncation variant in pancreatic lipase related protein 2 (PNLIPRP2) is expressed poorly and does not alter risk for chronic pancreatitis. PLoS One 13 : e0206869 . [ PMC free article ] [ PubMed ] [ Google Scholar ]
- Ollis D. L., Cheah E., Cygler M., Dijkstra B., Frolow F., Franken S. M., et al. (1992). The alpha/beta hydrolase fold. Protein Eng . 5 197–211. 10.1093/protein/5.3.197 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Pang W., Zhang Y., Wang S., Jia A., Dong W., Cai C., et al. (2011). The mPlrp2 and mClps genes are involved in the hydrolysis of retinyl esters in the mouse liver. J. Lipid Res. 52 934–941. 10.1194/jlr.M010082 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Payne R. M., Sims H. F., Jennens M. L., Lowe M. E. (1994). Rat pancreatic lipase and two related proteins: enzymatic properties and mRNA expression during development. Am. J. Physiol. 266 G914–G921. 10.1152/ajpgi.1994.266.5.G914 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Reboul E., Berton A., Moussa M., Kreuzer C., Crenon I., Borel P. (2006). Pancreatic lipase and pancreatic lipase-related protein 2, but not pancreatic lipase-related protein 1, hydrolyze retinyl palmitate in physiological conditions. Biochim. Biophys. Acta 1761 4–10. 10.1016/j.bbalip.2005.12.013 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Rippe C., Erlanson-Albertsson C., Lindqvist A. (2007). Consequences of metabolic challenges on hypothalamic colipase and PLRP2 mRNA in rats. Brain Res. 1185 152–157. 10.1016/j.brainres.2007.09.022 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Roussel A., de Caro J., Bezzine S., Gastinel L., de Caro A., Carrière F. (1998a). Reactivation of the totally inactive pancreatic lipase RP1 by structure-predicted point mutations. Proteins 32 523–531. [ PubMed ] [ Google Scholar ]
- Roussel A., Yang Y., Ferrato F., Verger R., Cambillau C., Lowe M. (1998b). Structure and activity of rat pancreatic lipase-related protein 2. J. Biol. Chem. 273 32121–32128. 10.1074/jbc.273.48.32121 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Sias B., Ferrato F., Pellicer-Rubio M. T., Forgerit Y., Guillouet P., Leboeuf B., et al. (2005). Cloning and seasonal secretion of the pancreatic lipase-related protein 2 present in goat seminal plasma. Biochim. Biophys. Acta. 1686 169–180. 10.1016/j.bbalip.2004.09.008 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Sodhi C. P., Fulton W. B., Good M., Vurma M., Das T., Lai C. S., et al. (2018). Fat composition in infant formula contributes to the severity of necrotising enterocolitis. Br. J. Nutr. 120 665–680. 10.1017/S0007114518001836 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Valek T., Kostelnik A., Valkova P., Pohanka M. (2019). Indoxyl Acetate as a Substrate for Analysis of Lipase Activity. Int. J. Anal. Chem. 2019 : 8538340 . 10.1155/2019/8538340 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- van Tilbeurgh H., Egloff M. P., Martinez C., Rugani N., Verger R., Cambillau C. (1993). Interfacial activation of the lipase-procolipase complex by mixed micelles revealed by X-ray crystallography. Nature 362 814–820. 10.1038/362814a0 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Wagner A. C., Wishart M. J., Mulders S. M., Blevins P. M., Andrews P. C., Lowe A. W., et al. (1994). GP-3, a newly characterized glycoprotein on the inner surface of the zymogen granule membrane, undergoes regulated secretion. J. Biol. Chem. 269 9099–9104. [ PubMed ] [ Google Scholar ]
- Winkler F. K., D’Arcy A., Hunziker W. (1990). Structure of human pancreatic lipase. Nature 343 771–774. 10.1038/343771a0 [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Withers-Martinez C., Carrière F., Verger R., Bourgeois D., Cambillau C. (1996). A pancreatic lipase with a phospholipase A1 activity: crystal structure of a chimeric pancreatic lipase-related protein 2 from guinea pig. Structure 4 1363–1374. [ PubMed ] [ Google Scholar ]
- Xiao X., Lowe M. E. (2015). The β5-Loop and Lid Domain Contribute to the Substrate Specificity of Pancreatic Lipase-related Protein 2 (PNLIPRP2). J. Biol. Chem. 290 28847–28856. 10.1074/jbc.M115.683375 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Xiao X., Mukherjee A., Ross L. E., Lowe M. E. (2011). Pancreatic lipase-related protein-2 (PLRP2) can contribute to dietary fat digestion in human newborns. J. Biol. Chem. 286 26353–26363. 10.1074/jbc.M111.249813 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Xiao X., Ross L. E., Sevilla W. A., Wang Y., Lowe M. E. (2013). Porcine pancreatic lipase related protein 2 has high triglyceride lipase activity in the absence of colipase. Biochim. Biophys. Acta 1831 1435–1441. 10.1016/j.bbalip.2013.06.002 [ PMC free article ] [ PubMed ] [ CrossRef ] [ Google Scholar ]
- Yang Y., Lowe M. E. (2000). The open lid mediates pancreatic lipase function. J. Lipid Res. 41 48–57. [ PubMed ] [ Google Scholar ]
- Yang Y., Sanchez D., Figarella C., Lowe M. E. (2000). Discoordinate expression of pancreatic lipase and two related proteins in the human fetal pancreas. Pediatr. Res. 47 184–188. 10.1203/00006450-200002000-00006 [ PubMed ] [ CrossRef ] [ Google Scholar ]

ENZYMES AND DIGESTION - The action of lipase on fats
Preparation of a series of different combinations.
- Do not use pipettes for any other purpose.
1) Place 6 test-tubes in a rack, and label them 1-6.
2) Using a single 3ml pipette (twice) put 6 ml of alkaline milk into tubes 1-6.
3) Using a single (smaller) pipette, add 0.5 ml phenolphthalein into tubes 1 to 5 BUT NOT 6.
4) Using a single different pipette, add 1 ml bile salts into tubes 1 , 2, 3 and 6 ONLY.
5) Using yet another pipette, add 1 ml lipase into tubes 1, 2, 4 and 6 ONLY.
6) Record the colours of the contents of the tubes in the table below - in the row labelled "colour at start".
7) Transfer tubes 2-6 to a water bath at 37 °C, and leave them for some time. In the meanwhile, you have the opportunity to carry out the test procedure overleaf which will tell you more about these substances.
8) At regular intervals examine the tubes and look for signs of colour changing from pink to whitish. Record the final colours of the contents of the tubes in the table below - in the row labelled "colour at end ".
| Tube number | 1 | 2 | 3 | 4 | 5 | 6 | |
|---|---|---|---|---|---|---|---|
| Contents | Quantity (ml) | ||||||
| | alkaline milk | 6 | 6 | 6 | 6 | 6 | 6 |
| phenolphthalein | 0.5 | 0.5 | 0.5 | 0.5 | 0.5 | - | |
| bile salts | 1 | 1 | 1 | - | - | 1 | |
| lipase | 1 | 1 | - | 1 | - | 1 | |
| / °C | lab | 37 | 37 | 37 | 37 | 37 | |
| colour at start | |||||||
| colour at end | |||||||
Which of the "contents "above is not normally found in the human small intestine?
> phenolphthalein
What is meant by the term lipolytic ?
> breaking down fats (lipids)into simpler substances
What is meant by the term emulsifying ?
> breaking (fat) droplets into smaller ones
Why do you think that the milk was made alkaline before the experiment? (2 reasons)
> because lipase acts in the alkaline conditions of the intestine > when the fatty acids are produced, they neutralise the alkali and cause the indicator to change colour
Whereabouts in the body is lipase produced, i.e. Which organ makes it?
> pancreas
Whereabouts in the body is lipase released, i.e Where does it mix with "food" to be digested?
> duodenum
What sort of physical conditions exist there?
> alkaline (warm, wet, etc)
Whereabouts in the body is bile produced, i.e. Which organ makes it?
Whereabouts in the body is bile stored?
> gall bladder
What do you think is the role of bile salts in this experiment?
> to emulsify fat, i.e. make droplets smaller and increase the surface area so as to make it more easily broken down by lipase
Which tube or tubes shows/show a change first?
What combination of ingredients and conditions do they include?
> lipase, bile salts, warm
Compare the result from this tube with the others. Write your conclusions from this experiment. This should include answers to questions such as:
What does lipase do to the fat in milk? What other factors are needed?Are they absolutely necessary?
> In order to break down fats into fatty acids, lipase needs warmth etc, and bile salts speed it up, but are not essential.
BIURET TEST
> protein
Pour about 10 ml of sodium (or potassium) hydroxide into a boiling tube.
CARE! Wear eye protection when handling alkalies.
Add about 1 ml of copper sulphate solution; the colour should change to a deeper blue. Mix carefully. This is biuret reagent.
Place about 2.5 ml of each substance to be tested into a tube in the rack.
To each add about 2.5 ml of the blue mixture ( biuret reagent ) produced above.
Wait for a different colour to develop, compared with the leftover untouched mixture.
| Test substance | Resulting colour | Conclusion : substances present |
|---|---|---|
| lipase | ||
| milk | ||
| amylase |
> The enzymes lipase and amylase are/contain protein - also milk is a protein
Finally: Pour away the contents of ALL the tubes down the sink, then rinse the tubes with a gentle flow of cold water before placing them in the washing up bowl.

- TOP CATEGORIES
- AS and A Level
- University Degree
- International Baccalaureate
- Uncategorised
- 5 Star Essays
- Study Tools
- Study Guides
- Meet the Team
- Molecules & Cells
The Action of Lipase and Bile Salts On Milk
THE ACTION OF LIPASE AND BILE SALTS ON MILK
To investigate the effect of temperature upon the action of lipase. Lipase is an enzyme that digests or breaks down fat into fatty acids and glycerol.
FACTORS INVOLVED
All factors will be kept constant with the exception of temperature that will be varied.
The factors to remain constant include:
- Concentration of the lipase enzyme
- Concentration of bile salts
- Volume of milk in each test tube
- Volume of phenol phthalein used
- Volume of NaCO 3
- Concentration of NaCO 3
A syringe will be used to put the bile salts into each test tube. The same syringe will be used for each test tube. This is to prevent cross-contamination as the syringe will only contain bile salts. Similarly a single syringe will be used to put the enzyme into each test tube. Again this is to prevent cross-contamination as this syringe will only contain the enzyme.
Immediately after the enzyme is added the solution will be stirred thoroughly. This is to encourage all of the enzyme to come into contact with the fat and the bile salts.
The temperature of the solution in each test tube will be regularly monitored by using a thermometer. The temperature of the water bath in each test tube will be regularly monitored by using a thermometer.
The lipase solution will be warmed to the temperature each water bath.
The thermometer will be cleaned after it is removed from each test tube.
A syringe will be used to put each of the components into the test tube. This enables the quantities used to be constant between the test tubes as it is accurate way of measuring.
Room Temperature
In a previous experiment it was demonstrated that the enzyme will operate at room temperature. The enzyme took 8 minutes to completely digest the fat. It is therefore predicted that the enzyme will operate at room temperature.

Increase in temperature
It is predicted that the rate of enzyme action will increase in proportion with the increase in temperature. The theory behind this prediction is known as the Kinetic Theory. This states that the higher the temperature is, the more collisions there will be between the substrate (fat) and the enzyme as a result of the increased energy of the enzyme. This means that as the temperature increases more enzyme will come into contact with more substrate and the reaction rate will therefore increase.
Optimum Rate
This is a preview of the whole essay.
The enzyme action will reach an optimum rate at a certain temperature.
Higher Temperatures
After this point the rate of enzyme action will slow despite the further rise in temperature. This is because the heat above a certain point changes the shape of the active site of the enzyme so that the substrate no longer fits. At a certain temperature the enzyme action will stop all together as the enzyme becomes denatured. This was demonstrated in a previous experiment where the enzyme amylase, which is used to break down starch into sugar, was placed in a starch solution at 35 degrees Celsius. The enzyme was shown to break down the starch into sugar. A sample of enzyme was then boiled and then placed in an identical starch solution. At the end of the experiment this solution was found to contain starch but no sugar. This is because the enzyme had become denatured and was unable to break the starch down into sugar.
The lipase used is synthetically produced and may therefore be more stable at higher temperatures than predicted. The optimum temperature may be higher than predicted and the temperature at which it becomes denatured may be higher than predicted.
Test tubes, test tube racks, water baths, beakers, thermometer, syringe, stop clock, stirring rod.
Laboratory rules were observed at all times.
Two test tubes will be placed in water baths at the following temperatures: 10 degrees; 20 degrees; 30 degrees; 60 degrees and 80 degrees. Two test tubes will be left at room temperature. The average time taken for each of the two tubes at each temperature will then be taken. Each test tube will be left in the water bath for 3 minutes.
In each of the 10 test tubes we added 3 cm 3 of milk, 3 cm 3 NaCO 3, 5 drops of phenolphthalein and 1 cm 3 bile salts. 1 cm 3 of lipase enzyme was placed in each of the water baths and left to reach the temperature of the water. After the lipase had been in the water baths for 3 minutes it was added to each of the test tubes. The two test tubes being tested at room temperature also had 1 cm 3 of lipase added. The time taken for the colour of the solution to change from pink to white was taken using a stop clock.
The results of the experiment are set out in the following table:
CONCLUSION & EVALUATION
At room temperature (22 degrees) the average time taken for the enzyme to digest all of the fat was 6 minutes 7 seconds.
At 10 degrees the average time taken for the enzyme to digest all of the fat was 14 minutes 3 seconds. This is therefore a decrease in the rate of enzyme action as the temperature decreases. This is in accordance with the prediction as enzyme action slows as temperatures decrease. This is because molecules move more slowly as temperature decreases so there will be fewer collisions between enzyme and substrate resulting in fewer reactions and a slower digestion time.
At 30 degrees the average time taken for the enzyme to digest all of the fat was 3 minutes 2 seconds. This is therefore a substantial increase in the rate of enzyme action from room temperature. This is in accordance with the prediction as enzyme action increases as temperatures increase. This is because molecules gain energy and move around more quickly resulting in more collisions and more reaction and a faster digression time.
At 60 degrees the average time taken for the enzyme to digest all of the fat was 24 minutes 2 seconds. This is therefore a substantial decrease in the rate of enzyme action from 30 degrees. This is again accordance with the prediction as enzyme action decreases as temperatures increase beyond a certain optimum temperature. The heat changes the shape of the enzyme so that the substrate no longer fits properly and the reaction slows.
At 80 degrees the enzyme did not digest any of the fat . The solution remained pink indefinitely. This because the shape of the enzyme had been changed by the heat so much that it was unable to bind with the substrate (the fat). The enzyme is said to be denatured. This is in accordance with the prediction as enzyme action was said to cease above a critical temperature. According to these results this critical temperature appears to be between 60 and 80 degrees.
According to these results the optimum temperature appears to be approximately 30 degrees as at this temperature the rate of enzyme action was fastest.
The bile salts were used to speed up the reaction. They physically break down the large fat molecules in the milk into smaller molecules which have a larger surface area so the lipase can chemically break down the fat molecules more easily.
The Phenol Phthalein was used because it is an indicator which is pink when alkaline and colourless when acidic. As a result of the presence of the NaCO 3 the solution is alkaline at the start of the experiment. The indicator is therefore pink. As the enzyme acts the fat is broken down into fatty acids and glycerol. The solution therefore becomes more acidic and the indicator becomes colourless. The rate of enzyme action can therefore be monitored according to the speed of change in the colour of the indicator.
The accuracy of the experiment may have been affected by the following factors:
Slight variations in the concentrations and volumes of enzyme, substrate and other substances used in each of the test tubes. This would be caused by inaccurate measurements of the substances.
Impurities in the solutions.
Temperatures may have fluctuated in test tubes as water baths not kept at a constant temperature.
To provide a more accuarate result the number of test tubes used in each temperature control could have been used. The average would then have been more reliable. More temperature controls could have been used (e.g. at 5 degree intervals). Natural lipase from the body could have been used in a separate experiment and the two experiments compared.

Document Details
- Word Count 1497
- Page Count 5
- Level AS and A Level
- Subject Science
Related Essays

The Comparison of Lipase Digestion with and without Bile Salts.

An investigation into the effect of lipase concentration on the rate of lip...

A2 coursework- The effects of bile salts on digestion of fat

Investigating The Activity Of The Enzyme Lipase On Milk
Advertisement
Effects of Surfactants on Lipase Structure, Activity, and Inhibition
- Expert Review
- Published: 14 January 2011
- Volume 28 , pages 1831–1842, ( 2011 )
Cite this article

- Vincent Delorme 1 , 2 ,
- Rabeb Dhouib 1 ,
- Stéphane Canaan 1 ,
- Frédéric Fotiadu 2 ,
- Frédéric Carrière 1 &
- Jean-François Cavalier 1
3652 Accesses
148 Citations
Explore all metrics
Lipase inhibitors are the main anti-obesity drugs prescribed these days, but the complexity of their mechanism of action is making it difficult to develop new molecules for this purpose. The efficacy of these drugs is known to depend closely on the physico-chemistry of the lipid-water interfaces involved and on the unconventional behavior of the lipases which are their target enzymes. The lipolysis reaction which occurs at an oil-water interface involves complex equilibria between adsorption-desorption processes, conformational changes and catalytic mechanisms. In this context, surfactants can induce significant changes in the partitioning of the enzyme and the inhibitor between the water phase and lipid-water interfaces. Surfactants can be found at the oil-water interface where they compete with lipases for adsorption, but also in solution in the form of micellar aggregates and monomers that may interact with hydrophobic parts of lipases in solution. These various interactions, combined with the emulsification and dispersion of insoluble substrates and inhibitors, can either promote or decrease the activity and the inhibition of lipases. Here, we review some examples of the various effects of surfactants on lipase structure, activity and inhibition, which show how complex the various equilibria involved in the lipolysis reaction tend to be.
This is a preview of subscription content, log in via an institution to check access.
Access this article
Subscribe and save.
- Get 10 units per month
- Download Article/Chapter or eBook
- 1 Unit = 1 Article or 1 Chapter
- Cancel anytime
Price includes VAT (Russian Federation)
Instant access to the full article PDF.
Rent this article via DeepDyve
Institutional subscriptions
Similar content being viewed by others
Effect of cyclic and acyclic surfactants on the activity of Candida rugosa lipase
Intrinsic lipolysis rate for systematic design of lipid-based formulations

Role of Surfactants and Their Applications in Structured Nanosized Systems
Abbreviations.
β-octyl glucoside
bovine serum albumin
critical micellar concentration
dog gastric lipase
electron paramagnetic resonance
diethyl p -nitrophenyl phosphate
guinea pig pancreatic lipase-related protein 2
human pancreatic lipase
sodium taurodeoxycholate
porcine pancreatic lipase
sodium dodecyl sulphate
site-directed spin labeling
triacylglycerol
tetraethylene glycol monooctyl ether
tetrahydrolipstatin (Orlistat)
Thermomyces lanuginosus lipase
Yarrowia lipolytica Lip2
Carrière F, Renou C, Ransac S, Lopez V, De Caro J, Ferrato F, et al . Inhibition of gastrointestinal lipolysis by Orlistat during digestion of test meals in healthy volunteers. Am J Physiol Gastrointest Liver Physiol. 2001;281(1):G16–28.
PubMed Google Scholar
Tiss A, Lengsfeld H, Verger R. A comparative kinetic study on human pancreatic and Thermomyces lanuginosa lipases: inhibitory effects of tetrahydrolipstatin in the presence of lipid substrates. J Mol Catal B Enzym. 2010;62:19–26.
Article CAS Google Scholar
Lengsfeld H, Beaumier-Gallon G, Chahinian H, De Caro A, Verger R, Laugier R, et al . Physiology of gastrointestinal lipolysis and therapeutical use of lipases and digestive lipase inhibitors. In: Müller G, Petry S, editors. Lipases and phospholipases in drug development. Weinheim: Wiley-VCH; 2004. p. 195–223.
Google Scholar
Ballinger A, Peikin S. Orlistat: its current status as an anti-obesity drug. Eur J Pharmacol. 2002;440(2–3):109–17.
Article PubMed CAS Google Scholar
Kopelman P, Bryson A, Hickling R, Rissanen A, Rossner S, Toubro S, et al . Cetilistat (ATL-962), a novel lipase inhibitor: a 12-week randomized, placebo-controlled study of weight reduction in obese patients. Int J Obes Lond. 2007;31(3):494–9.
Bryson A, de la Motte S, Dunk C. Reduction of dietary fat absorption by the novel gastrointestinal lipase inhibitor cetilistat in healthy volunteers. Br J Clin Pharmacol. 2009;67(3):309–15.
Yamada Y, Kato T, Ogino H, Ashina S, Kato K. Cetilistat (ATL-962), a novel pancreatic lipase inhibitor, ameliorates body weight gain and improves lipid profiles in rats. Horm Metab Res. 2008;40(8):539–43.
Kopelman P, Groot Gde H, Rissanen A, Rossner S, Toubro S, Palmer R, et al . Weight loss, HbA1c reduction, and tolerability of cetilistat in a randomized, placebo-controlled phase 2 trial in obese diabetics: comparison with orlistat (Xenical). Obesity (Silver Spring). 2010;18(1):108–15.
Tucci S, Boyland E, Halford J. The role of lipid and carbohydrate digestive enzyme inhibitors in the management of obesity: a review of current and emerging therapeutic agents. Diabetes Metab Syndr Obes. 2010;3:125–43.
Salem V, Bloom S. Approaches to the pharmacological treatment of obesity. Expert Rev Clin Pharmacol. 2010;3(1):73–88.
Article Google Scholar
Ben Ali Y, Chahinian H, Petry S, Muller G, Lebrun R, Verger R, et al . Use of an inhibitor to identify members of the hormone-sensitive lipase family. Biochemistry. 2006;45(47):14183–91.
Petry S, Baringhaus K-H, Schoenafinger K, Jung C, Kleine H, Müller G. High-throughput screening of hormone-sensitive lipase and subsequent. In: Müller G, Petry S, editors. Lipases and phospholipases in drug development. Weinheim: Wiley-VCH; 2004. p. 121–37.
Jin W, Millar JS, Broedl U, Glick JM, Rader DJ. Inhibition of endothelial lipase causes increased HDL cholesterol levels in vivo . J Clin Invest. 2003;111(3):357–62.
PubMed CAS Google Scholar
Cotes K, Bakala N'goma JC, Dhouib R, Douchet I, Maurin D, Carriere F, et al . Lipolytic enzymes in Mycobacterium tuberculosis. Appl Microbiol Biotechnol. 2008;78(5):741–9.
Mishra KC, de Chastellier C, Narayana Y, Bifani P, Brown AK, Besra GS, et al . Functional role of the PE domain and immunogenicity of the Mycobacterium tuberculosis triacylglycerol hydrolase LipY. Infect Immun. 2008;76(1):127–40.
Seibert G, Toti L, Wink J, inventors; Sanofi-Aventis, assignee. Use of cyclipostin derivatives for the treatment of mycobacterial infectious diseases. United States patent US 20090233884. 2009 Sep 17.
Dowling S, Cox J, Cenedella RJ. Inhibition of fatty acid synthase by Orlistat accelerates gastric tumor cell apoptosis in culture and increases survival rates in gastric tumor bearing mice in vivo . Lipids. 2009;44(6):489–98.
Jaeger K-E, Ransac S, Dijkstra BW, Colson C, Vanheuvel M, Misset O. Bacterial lipases. FEMS Microbiol Rev. 1994;15(1):29–63.
Gilbert EJ. Pseudomonas lipases: biochemical properties and molecular cloning. Enzyme Microb Technol. 1993;15(8):634–45.
Huang AHC. Plant lipases. In: Borgström B, Brockman HL, editors. Lipases. Amsterdam: Elsevier; 1984. p. 419–42.
Mukherjee KD, Hills MJ. Lipases from plants. In: Woolley P, Petersen SB, editors. Lipases: their structure, biochemistry and application. Cambridge: Cambridge University Press; 1994. p. 49–75.
Carrière F, Bezzine S, Verger R. Molecular evolution of the pancreatic lipase and two related enzymes towards different substrate selectivities. J Mol Catal B Enzym. 1997;3:55–64.
Carrière F, Gargouri Y, Moreau H, Ransac S, Rogalska E, Verger R. Gastric lipases: cellular, biochemical and kinetic aspects. In: Wooley P, Petersen SB, editors. Lipases: Their structure, biochemistry and application. Cambridge: Cambridge University Press; 1994. p. 181–205.
Alberghina L, Schmid RD, Verger R. Lipases: structure, mechanism, and genetic engineering. Weinheim: VCH; 1991.
Wooley P, Petersen SB. Lipases: their structure, biochemistry and applications. Cambridge: Cambridge University Press; 1994.
Brockerhoff H, Jensen RG. In: Brockerhoff H, Jensen RG, editors. Lipolytic enzymes. New York: Academic; 1974.
Borgström B, Erlanson C. In: Borgström B, Brockman HL, editors. Lipases. Amsterdam: Elsevier; 1984.
Aloulou A, Rodriguez JA, Fernandez S, Van Oosterhout D, Puccinelli D, Carriere F. Exploring the specific features of interfacial enzymology based on lipase studies. Biochim Biophys Acta, Mol Cell Biol Lipids. 2006;1761:995–1013.
CAS Google Scholar
Verger R, Mieras MCE, de Haas GH. Action of phospholipase A at interfaces. J Biol Chem. 1973;248(11):4023–34.
Panaitov I, Verger R. Enzymatic reactions at interfaces: interfacial and temporal organization of enzymatic lipolysis. In: Baszkin A, Norde W, editors. Physical chemistry of biological interfaces. New York: Marcel Dekker, Inc; 2000. p. 359–400.
Sarda L, Desnuelle P. Action de la lipase pancréatique sur les esters en émulsion. Biochim Biophys Acta. 1958;30:513–21.
Belle V, Fournel A, Woudstra M, Ranaldi S, Prieri F, Thome V, et al . Probing the opening of the pancreatic lipase lid using site-directed spin labeling and EPR spectroscopy. Biochemistry. 2007;46:2205–14.
Verger R. Interfacial enzyme kinetics of lipolysis. Annu Rev Biophys Bioeng. 1976;5:77–117.
Entressangles B, Desnuelle P. Action of pancreatic lipase on aggregated glyceride molecules in an isotropic system. Biochim Biophys Acta. 1968;159(2):285–95.
Antipova A, Semenova M, Belyakova L, Il’in M. On relationships between molecular structure, interaction and surface behavior in mixture: small-molecule surfactant+ protein. Colloids Surf B Biointerfaces. 2001;21(1–3):217–30.
Miller R, Fainerman V, Makievski A, Krägel J, Grigoriev D, Kazakov V, et al . Dynamics of protein and mixed protein/surfactant adsorption layers at the water/fluid interface. Adv Colloid Interface Sci. 2000;86(1–2):39–82.
Mogensen JE, Sehgal P, Otzen DE. Activation, inhibition, and destabilization of Thermomyces lanuginosus lipase by detergents. Biochemistry. 2005;44(5):1719–30.
Holmberg K, Jönsson B, Kronberg B, Lindman B. Surfactants and polymers in aqueous solution. Chischester: Wiley; 2002.
Book Google Scholar
Reynolds J, Tanford C. Binding of dodecyl sulfate to proteins at high binding ratios. Possible implications for the state of proteins in biological membranes. Proc Natl Acad Sci U S A. 1970;66(3):1002.
Otzen D, Oliveberg M. Burst-phase expansion of native protein prior to global unfolding in SDS. J Mol Biol. 2002;315(5):1231–40.
Otzen D. Protein unfolding in detergents: effect of micelle structure, ionic strength, pH, and temperature. Biophys J. 2002;83(4):2219–30.
Borgström B, Donner J. Interactions of pancreatic lipase with bile salts and dodecyl sulfate. J Lipid Res. 1976;17(5):491–7.
Brzozowski AM, Derewenda U, Derewenda ZS, Dodson GG, Lawson DM, Turkenburg JP, et al . A model for interfacial activation in lipases from the structure of a fungal lipase-inhibitor complex. Nature. 1991;351(6326):491–4.
Derewenda U, Brzozowski AM, Lawson DM, Derewenda ZS. Catalysis at the interface: the anatomy of a conformational change in a triglyceride lipase. Biochemistry. 1992;31(5):1532–41.
Brzozowski AM, Savage H, Verma CS, Turkenburg JP, Lawson DM, Svendsen A, et al . Structural origins of the interfacial activation in Thermomyces (Humicola) lanuginosa lipase. Biochemistry. 2000;39(49):15071–82.
Brzozowski AM. Crystallization of a Humicola lanuginosa lipase-inhibitor complex with the use of polyethylene glycol monomethyl ether. Acta Crystallogr D Biol Crystallogr. 1993;49(Pt 3):352–4.
Grochulski P, Bouthillier F, Kazlauskas RJ, Serreqi AN, Schrag JD, Ziomek E, et al . Analogs of reaction intermediates identify a unique substrate binding site in Candida rugosa lipase. Biochemistry. 1994;33(12):3494–500.
Egloff M-P, Marguet F, Buono G, Verger R, Cambillau C, van Tilbeurgh H. The 2.46 Å resolution structure of the pancreatic lipase-colipase complex inhibited by a C 11 alkyl phosphonate. Biochemistry. 1995;34(9):2751–62.
Roussel A, Canaan S, Egloff MP, Riviere M, Dupuis L, Verger R, et al . Crystal structure of human gastric lipase and model of lysosomal acid lipase, two lipolytic enzymes of medical interest. J Biol Chem. 1999;274(24):16995–7002.
Roussel A, Miled N, Berti-Dupuis L, Riviere M, Spinelli S, Berna P, et al . Crystal structure of the open form of dog gastric lipase in complex with a phosphonate inhibitor. J Biol Chem. 2002;277(3):2266–74.
van Tilbeurgh H, Egloff M-P, Martinez C, Rugani N, Verger R, Cambillau C. Interfacial activation of the lipase-procolipase complex by mixed micelles revealed by X-Ray crystallography. Nature. 1993;362(6423):814–20.
Article PubMed Google Scholar
Gonzalez-Navarro H, Bano MC, Abad C. The closed/open model for lipase activation. Addressing intermediate active forms of fungal enzymes by trapping of conformers in water-restricted environments. Biochemistry. 2001;40(10):3174–83.
Hermoso J, Pignol D, Kerfelec B, Crenon I, Chapus C, Fontecilla-Camps JC. Lipase activation by nonionic detergents. The crystal structure of the porcine lipase-colipase-tetraethylene glycol monooctyl ether complex. J Biol Chem. 1996;271:18007–16.
Miled N, Roussel A, Bussetta C, Berti-Dupuis L, Riviere M, Buono G, et al . Inhibition of dog and human gastric lipases by enantiomeric phosphonate inhibitors: a structure-activity study. Biochemistry. 2003;42(40):11587–93.
Rouard M, Sari H, Nurit S, Entressangles B, Desnuelle P. Inhibition of pancreatic lipase by mixed micelles of diethyl p -nitrophenyl phosphate and the bile salts. Biochim Biophys Acta. 1978;530:227–35.
Hermoso J, Pignol D, Penel S, Roth M, Chapus C, Fontecilla-Camps JC. Neutron crystallographic evidence of lipase-colipase complex activation by a micelle. EMBO J. 1997;16(18):5531–6.
Moreau H, Moulin A, Gargouri Y, Noël J-P, Verger R. Inactivation of gastric and pancreatic lipases by diethyl p -nitrophenyl phosphate. Biochemistry. 1991;30(4):1037–41.
Cudrey C, van Tilbeurgh H, Gargouri Y, Verger R. Inactivation of pancreatic lipases by amphiphilic reagents 5-(Dodecyldithio)-2-nitrobenzoic acid and tetrahydrolipstatin. Dependence upon partitioning between micellar and oil phases. Biochemistry. 1993;32(50):13800–8.
Tiss A, Miled N, Verger R, Gargouri Y, Abousalham A. Digestive lipases inhibition: an in vitro study. In: Müller G, editor. Lipases and phospholipases in drug development. Weinheim: Wiley-VCH; 2004. p. 155–93.
Ben Ali Y, Chahinian H, Petry S, Muller G, Carriere F, Verger R, et al . Might the kinetic behavior of hormone-sensitive lipase reflect the absence of the lid domain? Biochemistry. 2004;43(29):9298–306.
Desnuelle P, Sarda L, Ailhaud G. Inhibition de la lipase pancréatique par le diéthyl- p -nitrophényl phosphate en émulsion. Biochim Biophys Acta. 1960;37:570–1.
Gargouri Y, Chahinian H, Moreau H, Ransac S, Verger R. Inactivation of pancreatic and gastric lipases by THL and C12:0-TNB: a kinetic study with emulsified tributyrin. Biochim Biophys Acta. 1991;1085(3):322–8.
Hadvary P, Lengsfeld H, Wolfer H. Inhibition of pancreatic lipase in vitro by the covalent inhibitor tetrahydrolipstatin. Biochem J. 1988;256(2):357–61.
Borgström B. Mode of action of tetrahydrolipstatin: a derivative of the naturally occurring lipase inhibitor lipstatin. Biochim Biophys Acta. 1988;962(3):308–16.
Hogan S, Fleury A, Hadvary P, Lengsfeld H, Meier MK, Triscari J, et al . Studies on the antiobesity activity of tetrahydrolipstatin, a potent and selective inhibitor of pancreatic lipase. Int J Obes. 1987;11 Suppl 3:35–42.
Hochuli E, Kupfer E, Maurer R, Meister W, Mercadal Y, Schmidt K. Lipstatin, an inhibitor of pancreatic lipase, produced by Streptomyces toxytricini. II. Chemistry and structure elucidation. J Antibiot (Tokyo). 1987;40(8):1086–91.
Weibel EK, Hadvary P, Hochuli E, Kupfer E, Lengsfeld H. Lipstatin, an inhibitor of pancreatic lipase, produced by Streptomyces toxytricini. I. Producing organism, fermentation, isolation and biological activity. J Antibiot (Tokyo). 1987;40(8):1081–5.
Martinez C, Nicolas A, van Tilbeurgh H, Egloff M-P, Cudrey C, Verger R, et al . Cutinase, a lipolytic enzyme with a preformed oxyanion hole. Biochemistry. 1994;33(1):83–9.
Withers-Martinez C, Carriere F, Verger R, Bourgeois D, Cambillau C. A pancreatic lipase with a phospholipase A1 activity: crystal structure of a chimeric pancreatic lipase-related protein 2 from guinea pig. Structure. 1996;4(11):1363–74.
Lüthi-Peng Q, Winkler FK. Large spectral changes accompany the conformational transition of human pancreatic lipase induced by acylation with the inhibitor tetrahydrolipstatin. Eur J Biochem. 1992;205(1):383–90.
Tiss A, Carriere F, Verger R. Effects of gum arabic on lipase interfacial binding and activity. Anal Biochem. 2001;294(1):36–43.
Lookene A, Skottova N, Olivecrona G. Interactions of lipoprotein lipase with the active-site inhibitor tetrahydrolipstatin (Orlistat)(R). Eur J Biochem. 1994;222(2):395–403.
Tiss A, Lengsfeld H, Carrière F, Verger R. Inhibition of human pancreatic lipase by tetrahydrolipstatin: further kinetic studies showing its reversibility. J Mol Catal B Enzym. 2009;58(1–4):41–7. doi: 10.1016/j.molcatb.2008.11.003 .
Tiss A, Ransac S, Lengsfeld H, Hadvary P, Cagna A, Verger R. Surface behaviour of bile salts and tetrahydrolipstatin at air/water and oil/water interfaces. Chem Phys Lipids. 2001;111(1):73–85.
Tiss A, Lengsfeld H, Hadvary P, Cagna A, Verger R. Transfer of orlistat through oil-water interfaces. Chem Phys Lipids. 2002;119(1–2):41–9.
Gargouri Y, Piéroni G, Lowe PA, Sarda L, Verger R. Human gastric lipase. The effect of amphiphiles. Eur J Biochem. 1986;156:305–10.
Gargouri Y, Piéroni G, Rivière C, Saunière J-F, Lowe PA, Sarda L, et al . Kinetic assay of human gastric lipase on short- and long-chain triacylglycerol emulsions. Gastroenterology. 1986;91(4):919–25.
Bezzine S, Ferrato F, Ivanova MG, Lopez V, Verger R, Carriere F. Human pancreatic lipase: colipase dependence and interfacial binding of lid domain mutants. Biochemistry. 1999;38(17):5499–510.
Gargouri Y, Julien R, Bois AG, Verger R, Sarda L. Studies on the detergent inhibition of pancreatic lipase activity. J Lipid Res. 1983;24:1336–42.
Wahlgren M, Arnebrant T. Protein adsorption to solid surfaces. Trends Biotechnol. 1991;9(1):201–8.
Aloulou A, Puccinelli D, De Caro A, Leblond Y, Carriere F. A comparative study on two fungal lipases from Thermomyces lanuginosus and Yarrowia lipolytica shows the combined effects of detergents and pH on lipase adsorption and activity. Biochim Biophys Acta. 2007;1771(12):1446–56.
Chahinian H, Snabe T, Attias C, Fojan P, Petersen SB, Carriere F. How gastric lipase, an interfacial enzyme with a Ser-His-Asp catalytic triad, acts optimally at acidic pH. Biochemistry. 2006;45(3):993–1001.
Ranaldi S, Belle V, Woudstra M, Rodriguez J, Guigliarelli B, Sturgis J, et al . Lid opening and unfolding in human pancreatic lipase at low pH revealed by site-directed spin labeling EPR and FTIR spectroscopy. Biochemistry. 2009;48(3):630–8.
Gursoy RN, Benita S. Self-emulsifying drug delivery systems (SEDDS) for improved oral delivery of lipophilic drugs. Biomed Pharmacother. 2004;58(3):173–82.
Lipinski CA. Drug-like properties and the causes of poor solubility and poor permeability. J Pharmacol Toxicol Methods. 2000;44(1):235–49.
Charman W, Porter C, Mithani S, Dressman J. Physicochemical and physiological mechanisms for the effects of food on drug absorption: the role of lipids and pH. J Pharm Sci. 1997;86(3):269–82.
Pouton CW, Porter CJ. Formulation of lipid-based delivery systems for oral administration: materials, methods and strategies. Adv Drug Deliv Rev. 2008;60(6):625–37.
Charman SA, Charman WN, Rogge MC, Wilson TD, Dutko FJ, Pouton CW. Self-emulsifying drug delivery systems: formulation and biopharmaceutic evaluation of an investigational lipophilic compound. Pharm Res. 1992;9(1):87–93.
Attwood D, Florence AT. Surfactant systems: their chemistry, pharmacy and biology. London: Chapman and Hall; 1983.
Strickley R. Currently marketed oral lipid-based dosage forms: drug products and excipients. In: Hauss DJ, editor. Oral lipid-based formulations: enhancing the bioavailability of poorly water soluble drugs. New York: Informa Healthcare, Inc.; 2007. p. 1–31.
Fernandez S, Jannin V, Rodier JD, Ritter N, Mahler B, Carriere F. Comparative study on digestive lipase activities on the self emulsifying excipient Labrasol®, medium chain glycerides and PEG esters. Biochim Biophys Acta, Mol Cell Biol Lipids. 2007;1771:633–40.
Fernandez S, Rodier JD, Ritter N, Mahler B, Demarne F, Carrière F, et al . Lipolysis of the semi-solid self-emulsifying excipient Gelucire 44/14 by digestive lipases. Biochim Biophys Acta, Mol Cell Biol Lipids. 2008;1781:367–75.
Fernandez S, Chevrier S, Ritter N, Mahler B, Demarne F, Carriere F, et al . In vitro gastrointestinal lipolysis of four formulations of piroxicam and cinnarizine with the self emulsifying excipients Labrasol and Gelucire 44/14. Pharm Res. 2009;26(8):1901–10.
Christiansen A, Backensfeld T, Weitschies W. Effects of non-ionic surfactants on in vitro triglyceride digestion and their susceptibility to digestion by pancreatic enzymes. Eur J Pharm Sci. 2010;41(2):376–82.
Download references
ACKNOWLEDGMENTS
V. Delorme was financed by a PhD fellowship from the Ministère de l’Enseignement Supérieur et de la Recherche. This work was supported by the CNRS and by the Agence Nationale de la Recherche Française (ANR PCV 2007–184840 PHELIN, ANR MIEN 2009–00904 FOAMY_TUB). Authors would like to thank Dr. J. Blanc for revising the English.
Author information
Authors and affiliations.
CNRS - Aix-Marseille Université - Enzymologie Interfaciale et Physiologie de la Lipolyse, UPR 9025, 31 chemin Joseph Aiguier, 13402, Marseille cedex 20, France
Vincent Delorme, Rabeb Dhouib, Stéphane Canaan, Frédéric Carrière & Jean-François Cavalier
UMR 6263 CNRS-École Centrale Marseille-Université Paul Cézanne, Equipe Chirosciences, Avenue Escadrille Normandie-Niemen, 13397, Marseille Cedex 20, France
Vincent Delorme & Frédéric Fotiadu
You can also search for this author in PubMed Google Scholar
Corresponding authors
Correspondence to Frédéric Carrière or Jean-François Cavalier .
Rights and permissions
Reprints and permissions
About this article
Delorme, V., Dhouib, R., Canaan, S. et al. Effects of Surfactants on Lipase Structure, Activity, and Inhibition. Pharm Res 28 , 1831–1842 (2011). https://doi.org/10.1007/s11095-010-0362-9
Download citation
Received : 30 September 2010
Accepted : 27 December 2010
Published : 14 January 2011
Issue Date : August 2011
DOI : https://doi.org/10.1007/s11095-010-0362-9
Share this article
Anyone you share the following link with will be able to read this content:
Sorry, a shareable link is not currently available for this article.
Provided by the Springer Nature SharedIt content-sharing initiative
- interfacial enzymology
- lipid-protein interaction
- Find a journal
- Publish with us
- Track your research

IMAGES
VIDEO
COMMENTS
Investigating effect of temperature on the activity of lipase
Another great extension exercise is to test how an absence of bile salts affects the experiment. To do this, place Lipase in both test tubes, but replace the bile salts with the same volume of water in the control test tube. Bile salts reduce the particle size of the fat droplets which increases the surface area, thereby allowing better access ...
meaning that gastric lipase does not work at its optimum pH in the stomach (although pepsin - a protease that has an optimum pH of 1.5 to 2 is closer to its optimum pH). However, unlike pancreatic lipase, gastric lipase does not require bile salts to emulsify the fats first and is not denatured by the extremely acidic conditions of the stomach.
In today's experiment, you will investigate the digestion of fats by pancreatic lipase in the presence and absence of bile salts. Half and half will be used as the source of fat; it will be premixed with litmus powder, a chemical that is blue/purple in alkaline environments and pink in acidic environments.
Bile salts are slightly alkaline, with pH range of about 7-8 (Britannica). This helps lipase in catalysing its reaction. Bile salts also help lipase by increasing the surface area of fat droplets. Bile molecules have a hydrophobic and hydrophilic part. The hydrophobic part is attracted towards fat while the hydrophilic part is attracted towards ...
To do this, place Lipase in both test tubes, but replace the bile salts with the same volume of water in the control test tube. Bile salts reduce the particle size of the fat droplets which increases the surface area, thereby allowing better access for the Lipase to break down the triglycerides faster. Discuss the use of bile in the digestive ...
Bile salts affect the equilibrium reaction (Fig. 5) in such a way that at low pH values the amount of monoglycerides increases and that of triglycerides decreases. At alkaline pH the bile salt effect is different using rat and human pancreatic lipase (see Fig. 3 and 5).
This effect is best explained The recently isolated "colipase" (10) invites some speculations. We find that the protein, albumin, protects lipase (Fig. 4) and. by the hypothesis of Benzonana (5) that long chain acids at the interphase inhibit the lipase but are removed by complex formation with bile salts.
pancreatic lipase. However, the presence of bile salts, necessary to remove lipolysis products andpolar lipids from theinterface, also inhibit pancreatic lipase activity.7 Therefore, pancreatic lipase requires the cofactor colipase in order to act effectively in the presence of bile salts.7,8 Colipase has a molecular mass about 10 kDa with 93 ...
Bile Salt-Stimulated Lipase and Pancreatic ...
Bile salt-stimulated lipase (BSSL) is another important enzyme involved in digestion and absorption of dietary fat. ... Stromqvist M, Hernell O. Recombinant human milk bile salt-stimulated lipase. Catalytic activity is retained in the absence of glycosylation and the unique proline-rich repeats. J Biol Chem. 1993; 268:26692-26698. [Google ...
Bile Salt-Dependent Lipase - an overview
Structure and Function of Pancreatic Lipase-Related ...
The enzyme lipase is from the mammalian digestive system. Depending on the conditions in the surrounding medium, it may break down (digest) the fat to fatty acids and glycerol. During this practical session you will be seeing the lipolytic effect of the enzyme (as well as the contribution made by the emulsifying effects of bile salts).
Bile Salts and Lipase Experiment By: Grace and Jamie Purpose Purpose What is the effect of adding bile salts and altering temperature in the enzymatic action of lipase? Hypothesis Hypothesis Test Tube A Test Tube A Test tube A will stay the same because there is no lipase added, the.
stimulated lipase (BSSL) and bile salt-stimulated esterase (BSSE) activities in colostrum were similar to corresponding enzyme activities in transitional milk and in mature milk.
Bile salts affect the equilibrium reaction (Fig. 5) in such a way that at low pH values the amount of mono- glycerides increases and that of triglycerides decreases. At alkaline pH the bile salt effect is different using rat and human pancreatic lipase (see Fig. 3 and 5).
Similarly a single syringe will be used to put the enzyme into each test tube. Again this is to prevent cross-contamination as this syringe will only contain the enzyme. Immediately after the enzyme is added the solution will be stirred thoroughly. This is to encourage all of the enzyme to come into contact with the fat and the bile salts.
In this context, surfactants have important effects on lipase activity and inhibition, and the aim of this paper is to review some of the specific effects of surfactants, including synthetic compounds, bile salts, proteins and phospholipids. Lipases occur widely in the microbial (14, 18, 19), plant (20, 21) and animal kingdoms (22, 23).
Biology, AQA, GCSE, cyberteachers, cyber teachers, protein, enzymes, enzyme, how do enzymes work, biological catalyst, biological catalysts, active site, sub...